Simple features now on CRAN
Submitting a package to CRAN is always an adventure, submitting a package with lots of external dependencies even more so. A week ago I submitted the simple features for R package to CRAN, and indeed, hell broke loose! Luckily, the people behind CRAN are extremely patient, friendly and helpful, but they let your code test on a big server farm with computers in 13 different flavors.
Of course we test code on linux and windows after every code push to github, but that feels like talking to a machine, like remotely compiling and testing. CRAN feels different: you first need to manually confirm that you did your utmost best to solve problems, and then there is a person telling you everything still remaining! Of course, this is of incredible help, and a big factor in the R community’s sustainability.
Package sf is somewhat special in that it links to GEOS and GDAL, and in particular GDAL links, depending on how it is installed, to many (77 in my case) other libraries, each with their own versions. After first submission of sf 0.2-0, I ran into the following issues with my code.
sf 0.2-0
- I had to change all links starting with
http://cran.r-project.org/web/packages/pkgintohttps://cran.r-project.org/packages=pkg. A direct link to a units vignette on CRAN had to be removed. - some of the tests gave very different output, because my default testing platforms (laptop, travis) have PostGIS, and CRAN machines don’t; I changed this so that testing without PostGIS (CRAN) is now most silent
- the tests still output differences in GDAL and GEOS versions, but that was considered OK.
That was it! The good message
Thanks, on CRAN now.
Best
-k
arrived! Party time! Too early. In the evening (CRAN never sleeps) an email arrived, mentioning:
This worked for my incoming checks, but just failed for the regular
checks, with
Error in loadNamespace(name) : there is no package called 'roxygen2'
Calls: :: ... tryCatch -> tryCatchList -> tryCatchOne -> <Anonymous>
Execution halted
For some reason roxygen2 is not working.
Is it installed?
ERROR: configuration failed for package ‘sf’
with lots of helpful hints. Indeed, my package generated Rcpp files and manual pages dynamically during install; this requires Rcpp and roxygen2 to be available unconditionally and they aren’t.
So I sat down and worked on 0.2-1, to address this. Before I could do that, an email from Oxford (Brian Ripley) arrived, telling me that sf had caused some excitement in the multi-flavor server farm:
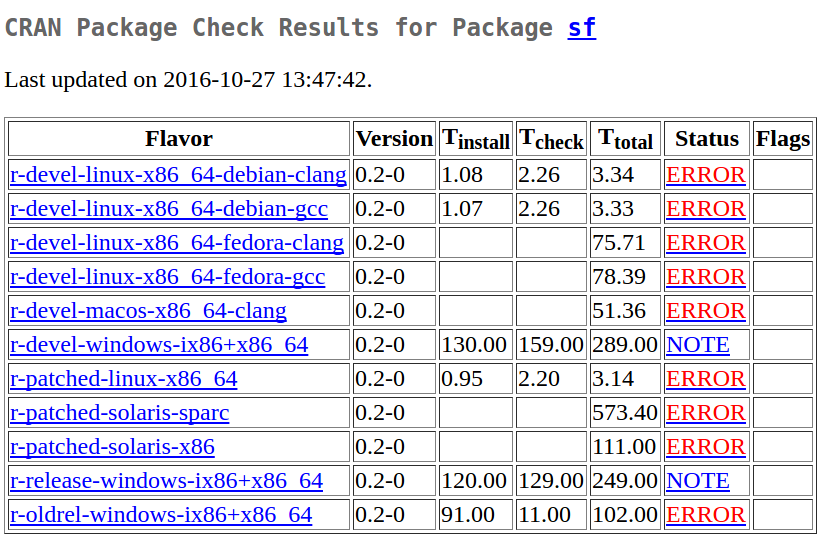
Here, it should be noted (again) that the only two NOTEs were due to the excellent work of Jeroen Ooms who compiled GDAL and many other libraries for rwinlib, and prepared sf for downloading and using them. The rest was my contribution.
In addition, an issue was raised by Dirk Eddelbuettel, telling me that his Rcpp reverse check farm had noted him that sf required GDAL 2.0 or later, but not by properly checking its version but by generating plane compile errors. The horror, the horror.
sf 0.2-1: roxygen, Rcpp, SQLITE on Solaris
sf 0.2-1 tried to address the Rcpp and roxygen2 problems: I took
their generation out of the configure and configure.win scripts.
I added all automatically derived files to the github repo, to get
everything in sync. Worked:
Thanks, on CRAN now. [Tonight I'll know for sure ...]
Best
-k
… and no emails in the evening.
Also, the errors on Solaris platforms were caused by the SQLITE library not being present, hence GeoPackage not being available as a GDAL driver. As a consequence, I had to set back the examples reading a GeoPackage polygons file to one where a shapefile is read. Bummer.
Another issue was related to relying on GDAL 2.1 features without testing for it; this was rather easily solved by conditional compiling.
This gave:
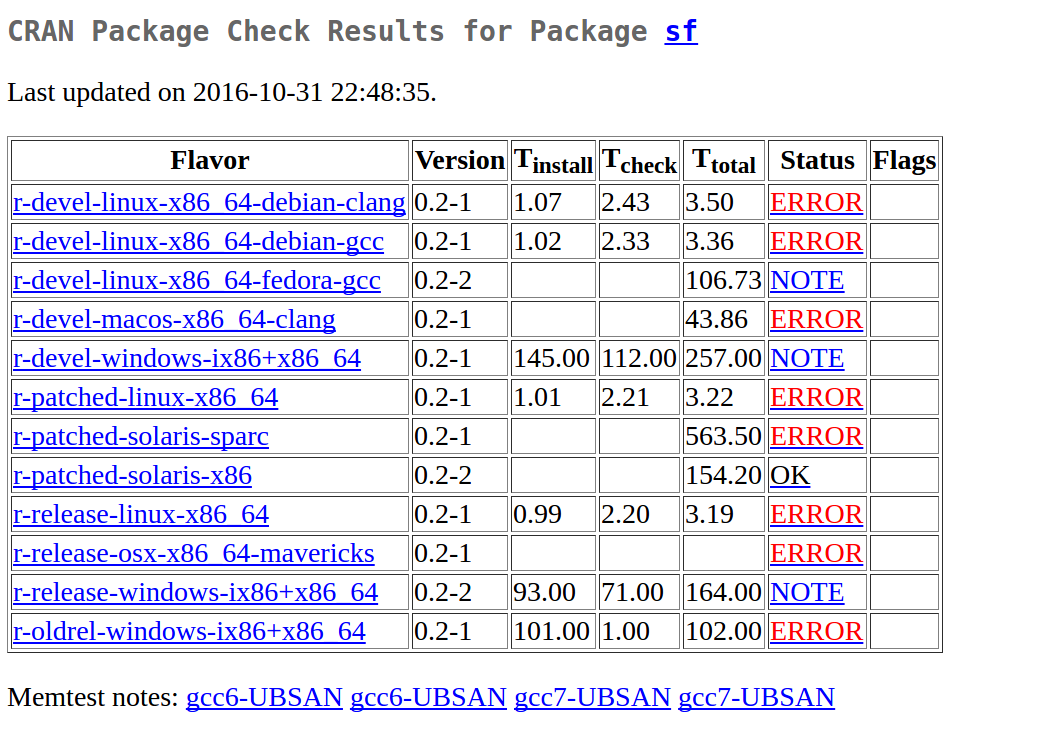
meaning SOME improvement, but where do these UBSAN reports suddenly come from!?
sf 0.2-2: byte swapping, memory alignment
Over-optimistically as I am, I had commented that life is too short to do byte swapping when reading WKB. Welcome to the CRAN server farm. Solaris-Spark is big-endian. Although the R code reading WKB does read non-native endian, this would have required to rewrite all tests, so I added byte swapping to the C++ code, using a helpful SO post.
The UBSAN issues all listed something like:
UBSAN: don't assume that pointers can point anywhere to, in valid memory;
wkb.cpp:185:39: runtime error: load of misaligned address 0x6150001d0e29 for type 'uint32_t', which requires 4 byte alignment
0x6150001d0e29: note: pointer points here
00 00 00 01 06 00 00 00 01 00 00 00 01 03 00 00 00 01 00 00 00 1b 00 00 00 00 00 00 a0 41 5e 54
Sounds scary? What I had done was for every coordinate use a double pointer, pointing it to the right place in the WKB byte stream, then copy its value, and move it 8 bytes. I love this *d++ expression. But you can’t do this anymore! Although the code worked on my machines, you can’t put a double pointer to any location and assume it’ll work everywhere. The solution was to memcpy the relevant bytes to a double value on the stack, and copy that into the Rcpp::NumericVector.
To be done:
All these changes have brought me here:
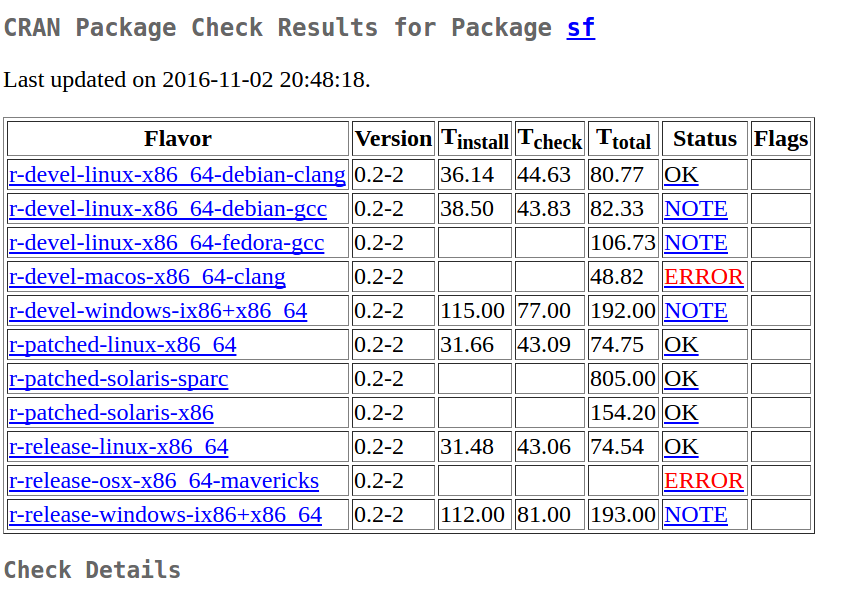
where you see that linux and Windows compile (all NOTEs indicate that the library is too large, which is unavoidable with GDAL) and that errors are Mac related:
r-devel-macos-x86_64-clang
** testing if installed package can be loaded
Error in dyn.load(file, DLLpath = DLLpath, ...) :
unable to load shared object '/Users/ripley/R/packages/tests-devel/sf.Rcheck/sf/libs/sf.so':
dlopen(/Users/ripley/R/packages/tests-devel/sf.Rcheck/sf/libs/sf.so, 6): Symbol not found: _H5T_NATIVE_DOUBLE_g
Referenced from: /Users/ripley/R/packages/tests-devel/sf.Rcheck/sf/libs/sf.so
Expected in: flat namespace
in /Users/ripley/R/packages/tests-devel/sf.Rcheck/sf/libs/sf.so
which indicates a problem with GDAL not linking to the HDF5 library (unsolved).
The second:
r-release-osx-x86_64-mavericks
checking gdal-config usability... yes
configure: GDAL: 1.11.4
checking GDAL version >= 2.0.0... yes
indicates that
- this platform still runs GDAL 1.x, so needs to be upgraded and that
- my check for GDAL version present on the system still does not work!
CRAN flavors
CRAN flavors is a great asset that teaches you problems of all kinds at an early stage. Without it, users would have run at some stage into problems that are now caught up front. Thanks to the tremendous effort of the CRAN team!!
