Experimenting with the R-ArcGIS bridge
Around a year ago, ESRI, market leader in commercial geographic information systems, announced the R-ArcGIS bridge, a software development meant to (quote) ``[c]ombine the power of ArcGIS and R to solve your spatial problems.’’
A MSc student in my group, Shankarlingam Sundaresan, worked for five months on a student assistant contract (10 hrs/wk) to develop some simple spatial statistical applications that could demonstrate the use of this bridge, and the possibilities and challenges of integrating functionality from some R packages in ArcGIS. We focused on package gstat for geostatistics, and package spatstat for point pattern analysis.
The result of Shankar’s work is found in his github repo which I forked here. Example runs are documented for
As you can see from the R code, much of the code is really administration: getting the right variables out of the ArcGIS database, and putting the right outputs again back in. Some of the work is of course not in the R code, but in code that defines the ArcGIS plugin and user interface. This interface then calls, with help of the arcgisbindings R package, the code in this package.
An interesting issue we found is that currently, the R-ArcGIS bridge does not support raster data. For both geostatistics and point pattern analysis, rasters are pretty essential, either for interpolating values on a regular grid, or estimating point pattern densities. The trick to ``show’’ raster data returned by R in ArcGIS (inspired by one of the examples from ESRI) is to return them as points, and display them as small coloured squares, as in:
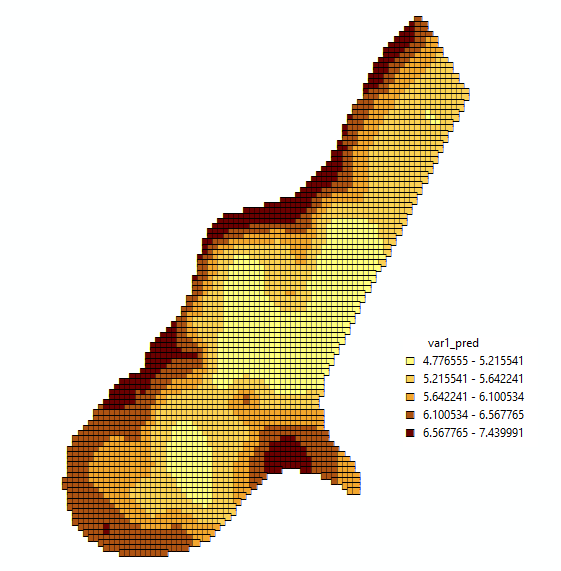
You can still see the at the right an bottom sides. Resizing the figure destroys the pattern completely, and would call for a different symbol size! Another possibility is to classify grid cells and merge them into polygons, which is done by the point pattern example. This looks better:
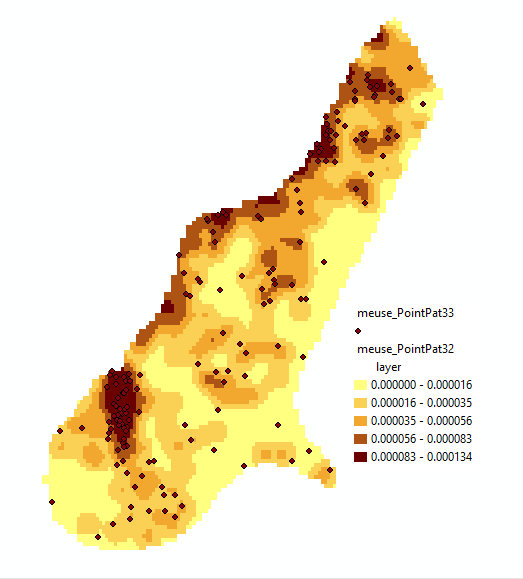
but of course looses the individual grid cell values, meaning you can not modify class boundaries later on. Regrettably, the time failed to do tests with lines data, or with methods that work on areas (spatial regression models, or disease models).
We further ran in some challenges to reproduce an error raised by
ArcGIS, and in the inflexibility of user interfaces: you can program
it such that it lets you choose a variable in the attribute table,
but if you do that you can’t transform it, say, by taking its
logarithm. If you want that flexibility, say, define a variable
by a formula such as log(zinc)~1, then you need to define it as
a character string input and loose the possibility to pick the
variable (zinc) from a drop-down list.
Who will use this bridge?
Writing these interfaces, elementary as they are, was quite a bit of work, which of course gets faster when you’ve done it a couple of times. None of this work adds something to R, or to the R packages used, in terms of functionality for R users. The intended audience for such interfaces is clear: users who want R functionality without having to leave ArcGIS (meaning: without having to understand R).
Who will develop such interfaces? Users who already use R probably know it is generally better to develop an R script than use an interactive interface, because in contrast to interactive work, the script documents what you did and lets you share and reproduce it.
Many R package developers start off programming because it solves their own problems, and share code when they feel it might solve other people’s problems too. They maintain code because the interaction with users asking questions, raising issues or providing contributions may be a nice way to get in touch with people, and learn from them. Programmers for the R-ArcGIS bridge interfaces will program primarily to solve other people’s problems, and will have to understand very well the R packages they interface, as well as understand how ArcGIS users want to interact with their software. What will motivate them to do this? Where will users R through the R-ArcGIS bridge have to go with their questions?
Tom Hengl, a good friend of mine, visited the 90 minutes short course on How to build tools for spatial-temporal modelling with R and ArcGIS, which was held by Ionut Cosmin Sandric during EGU 2016. He reported that between 60 and 100 people attended it, in a conference of 13,650 geoscientists – not bad!
Can you do this legally?
Whenever two softwares are closely linked, one commercial and closed source and the other largely open source under the GPL, one should wonder to what extent the development is legal.
R however is not completely under the GPL: It has two
ways of interfacing: one is via the R packaging mechanism and
Rinternals.h
and falls under the LGPL, the
other is for instance through functions in
Rembedded.h,
and falls under the GPL. The LGPL allows linking to non-GPL code,
the GPL requires linking to GPL (or GPL-compatible) code.
I looked into the source files, and didn’t notice any GPL header included, or function declared in a GPL header called, so that is good; it means that ArcGIS cannot start and stop R, and thus completely hide it as a back-end.
I didn’t check what is exactly linked against what, since the makefiles assumes (Microsoft) compilers I don’t have access to. The R-ArcGIS bridge github repo contains a large binary object, which seems to be a binary library for which source code and any other information are missing. Strange.
What about joining the R consortium?
Although ESRI could also have opted to bridge to Tibco’s TERR, which was designed for commercial embedding, it chose to link to R which is free and open source. This definitely makes it cheaper for new users to adopt it.
Not surprisingly, ESRI is hot on doing open source projects as long as ArcGIS is on one side of the equation. On the other side of this equation, however, people are developing and maintaining R usually without earning money or getting profits. The R consortium was recently founded to mediate between a group of enterprises benefiting from R and the open source communities developing and maintaining R. It would show good open source citizenship if ESRI would also join this consortium.
